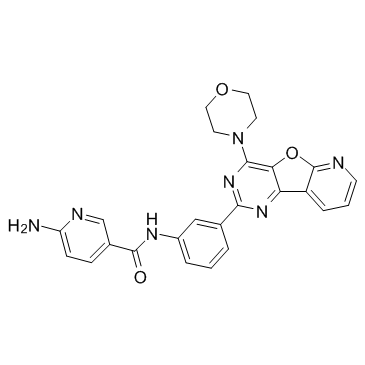
371942-69-7
| Name | 6-amino-N-[3-(4-morpholin-4-ylpyrido[2,3]furo[2,4-b]pyrimidin-2-yl)phenyl]pyridine-3-carboxamide |
|---|---|
| Synonyms |
3-Pyridinecarboxamide, 6-amino-N-[3-[4-(4-morpholinyl)pyrido[3',2':4,5]furo[3,2-d]pyrimidin-2-yl]phenyl]-
Unable to generate chemical name cc-386 6-Amino-N-{3-[4-(4-morpholinyl)pyrido[3',2':4,5]furo[3,2-d]pyrimidin-2-yl]phenyl}nicotinamide YM-201636 |
| Description | YM-201636 is a potent and selective PIKfyve inhibitor with an IC50 of 33 nM. YM-201636 also inhibits p110α with IC50 of 3.3 μM. |
|---|---|
| Related Catalog | |
| Target |
PIKfyve:33 nM (IC50) p110α:3.3 μM (IC50) Autophagy |
| In Vitro | Acute treatment of cells with YM-201636 shows that the PIKfyve pathway is involved in the sorting of endosomal transport, with inhibition leading to the accumulation of a late endosomal compartment and blockade of retroviral exit. The yeast orthologue of PIKfyve, Fab1, is found to be insensitive to YM-201636 (IC50>5 μM). YM-201636 does not inhibit a type IIγ PtdInsP kinase even at 10 μM and inhibits a mouse type Iα PtdInsP kinase with an IC50>2 μM[1]. YM-201636 almost completely inhibits basal and insulin-activated 2-deoxyglucose uptake at doses as low as 160 nM, with IC50=54 nM for the net insulin response. YM-201636 also completely inhibits insulin-dependent activation of class IA PI 3-kinase[2]. At low doses (10-25 nM), YM-201636 inhibits preferentially PtdIns5P rather than PtdIns(3,5)P2 production, whereas at higher doses, the two products are similarly inhibited. YM-201636 at 160 nM inhibits PtdIns5P synthesis twice more effectively compared with PtdIns(3,5)P2 synthesis[3]. MDCK cells treated with YM-201636 accumulate the tight junction protein claudin-1 intracellularly. YM-201636 treatment blocks the continuous recycling of claudin-1/claudin-2 and delays epithelial barrier formation[4]. |
| Kinase Assay | Following 3T3L1 adipocyte serum-starvation and insulin stimulation, cell lysates containing protease inhibitors are clarified and then subjected to immunoprecipitation with anti-PIKfyve antibodies. Washed beads are mixed with 100 μM PtdIns and preincubated for 15 min with YM-201636 (100 nM) or vehicle in the assay buffer (50 mM Tris-HCl, pH 7.5, 1 mM EGTA and 10 mM MgCl2). The kinase assay (50 μL final volume) is carried out for 15 min at 37 °C with 15 μM ATP and [γ-32P]ATP (30 μCi). Lipids are extracted, spotted on TLC glass plates (250 μm), resolved by a chloroform/methanol/water/ammonia solvent system and detected by autoradiography[2]. |
| Cell Assay | YM-201636 is dissolved in DMSO and diluted with DMEM and added to cells at a final concentration of 800 nM. Cells are treated with YM-201636 or a DMSO control for 2 h. For TER measurements cells are plated at confluency on Transwell permeable polyester filters (0.4 µm pore size) with surface area of 0.33 cm2. Media is changed ever 2-3 days and cells are grown for 7 days prior to TER measurements[4]. |
| References |
| Density | 1.4±0.1 g/cm3 |
|---|---|
| Molecular Formula | C25H21N7O3 |
| Molecular Weight | 467.479 |
| Exact Mass | 467.170593 |
| PSA | 132.29000 |
| LogP | 2.05 |
| Appearance | light yellow solid |
| Index of Refraction | 1.751 |
| Storage condition | -20℃ |